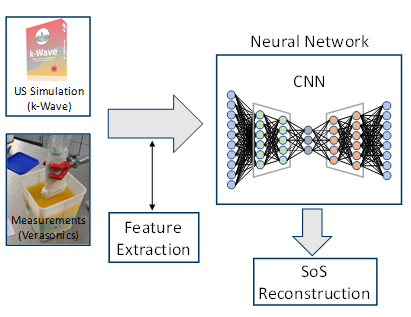
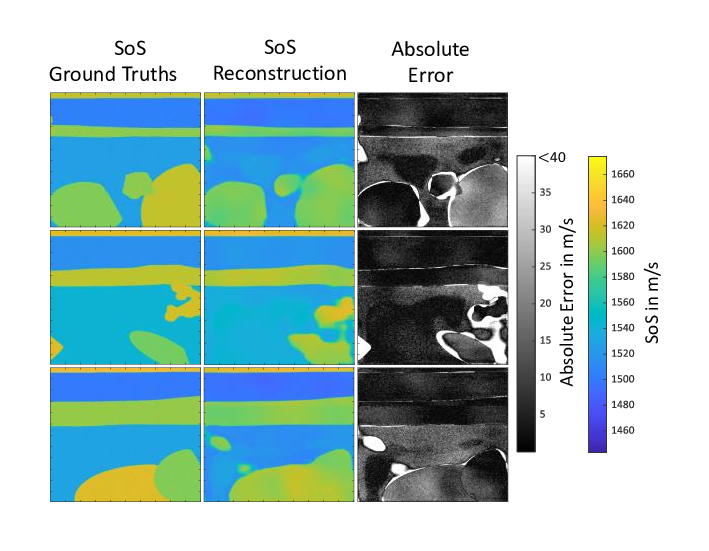
The quantitative reconstruction of material parameters using tomographic reconstruction methods such as the so-called Contrast Source Inversion or the Kaczmarz method are very computationally intensive and time-consuming. The high time expenditure is a consequence of the fact that an inverse problem has to be solved, which on the one hand represents a so-called ill-posed as well as non-linear problem. To solve this inverse problem, a time-consuming iterative data fitting procedure is used. As a novel, alternative reconstruction method, work is currently being done on new deep-learning-based methods. A so-called Convolutional Neural Network is trained on simulated ultrasound data, specifically on a coherence measure derived from the data, together with the ground truths to be reconstructed in the form of the speed of sound (SoS) distribution.
The aim is to enable the neural network to learn the non-linear relationship between measured ultrasound data and the material parameter distribution in the form of the SoS. Instead of using ultrasonic measurements with ring-shaped transducers setups enclosing the medium to be imaged, which are necessary for classical tomographic reconstruction methods, this method uses conventional linear ultrasonic transducers. The challenges of this approach are to generate a sufficiently large and realistic training data set and to transfer the technique from the application on simulation data to the application on real ultrasound data.
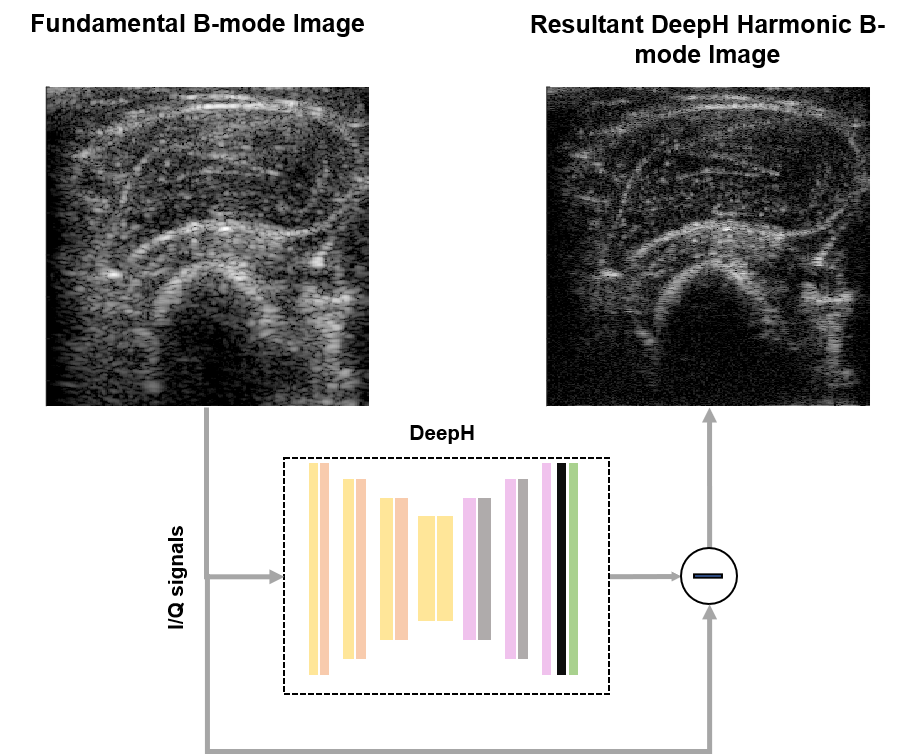
Tissue Harmonic Imaging (THI) is an invaluable tool in clinical ultrasound owing to its enhanced contrast resolution and reduced reverberation clutter in comparison to fundamental mode imaging. However, harmonic content separation based on high pass filtering suffers from potential contrast degradation or lower axial resolution due to spectral leakage. Whereas nonlinear multi-pulse harmonic imaging schemes, such as amplitude modulation and pulse inversion, suffer from a reduced framerate and comparatively higher motion-artifacts due to the necessity of at least two pulse echo acquisitions. To address this problem, we propose a deep-learning-based single-shot harmonic imaging technique capable of generating comparable image quality to conventional methods, yet at a higher framerate and with fewer motion artefacts. The model was evaluated across various targets and samples to illustrate generalizability as well as the possibility and impact of transfer learning. Hence, we demonstrate the ability of the proposed approach to generate harmonic images with a single firing that are comparable to those from a multi-pulse acquisition and outperform those acquired by linear filtering.
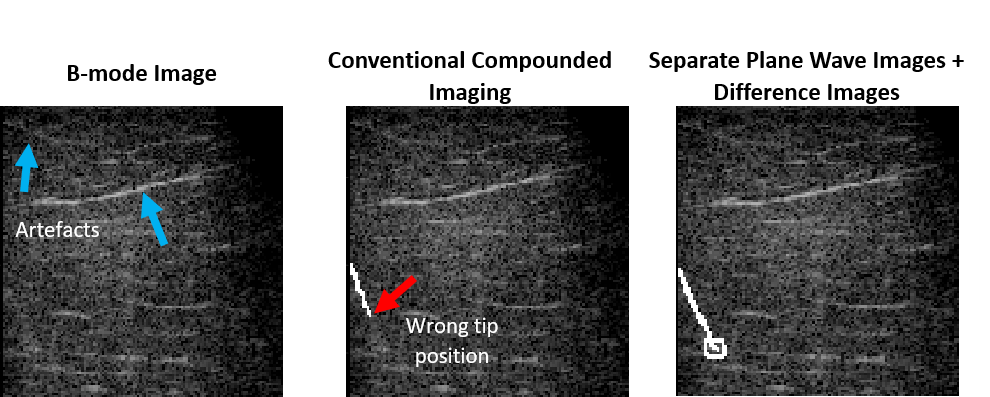
Sonography is commonly used to monitor the insertion of a needle into human tissue. However, good visibility in B-mode images is rarely guaranteed, especially with deep punctures and steep insertion angles. We hypothesize that the utilization of deep neural networks is particularly suitable to process this nonlinear needle-related information. Moreover, we presume that the exploitation of different information found in different plane waves could enhance the localization of the needle compared to the direct utilization of compounded B-mode images. Therefore, a deep learning-based framework capable of extracting the information provided by the separate plane waves is employed to enhance the localization of the shaft and tip of the needle. Using separate plane wave B-mode images and their difference images substantially improved the visualization of the needle by achieving 4.5-times lower tip localization error and improving the estimation of the puncture angle by 30.6% compared to the direct utilization of compounded B-mode images.